Présentation
To design effective ways to mitigate the spread of a pathogen in a population, it is important to first have a good understanding of the transmission characteristics of the pathogen as well as the risk factors for transmission. Ideally, such assessment should be performed from data documenting precisely the chains of transmission (i.e. informing on the source of infection of each case). Unfortunately, apart from few exceptions, such data are rarely available. While we may relatively easily document who got sick and when they got sick, it is much harder to identify who was infected by whom. This “missing data” problem puts us in a difficult position: we never observe any transmission even; how then can we evaluate transmission?
The Mathematical Modelling of Infectious Diseases Unit is specialized in the development of statistical and mathematical techniques to tackle this complex challenge. The kind of method we use will depend on the nature of the data and the research question.
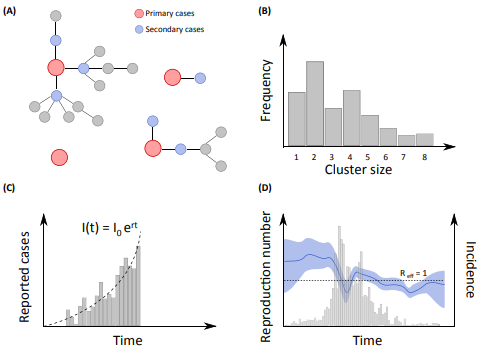
In situations where detailed epidemiological investigations have been performed to identify cases in a community, we often rely on Bayesian data augmentation techniques to probabilistically reconstruct unobserved chains of transmission and estimate key transmission parameters. These approaches are computationally intensive but extremely powerful to tackle complex missing data issues. We have used them to address a number of important epidemiological questions such as identifying the determinants of influenza transmission in households, characterizing how social networks structure the spread of influenza in schools or how geography and behaviors affect arbovirus transmission in small communities.
Often, however, we only have access to more aggregated surveillance data (e.g. daily counts of cases). In these situations, we can use simple methods that assess the transmission potential and the impact of control measures from the epidemic growth rate or more sophisticated mechanistic approaches that explicitly model the underlying transmission process, often with ordinary differential equations or their stochastic equivalent. We then rely on a multitude of techniques for the estimation of model parameters (e.g. MCMC, pMCMC, ABC, particle filters).
Other types of data we often work with include transversal or longitudinal serological studies, data documenting clusters of cases for zoonotic diseases and more recently genotypic viral data, social media data and mobile phone data. A challenging but exciting part of our research is that almost each outbreak is different from the other ones and comes with its own set of quite specific data and questions so that we always have to be inventive and develop new ways to make the most out of the data we get.