Présentation
Cell-cell signaling mediated by Notch receptors regulates a wide range of developmental processes and perturbations of Notch signaling activity underlie various human developmental and adult-onset diseases, including T-cell leukemia and other cancers. Additionally, Notch controls homeostasis in several adult tissues (skin, intestine, blood vessels) by regulating the quiescence, division and differentiation of adult stem cells. Thus, a molecular understanding of the control mechanisms of Notch signaling is of both fundamental and practical importance in molecular medicine.
Notch receptors aretransmembrane proteins with an extracellular ligand-binding domain and an intracellular domain acting as a membrane-tethered transcriptional activator. Notch receptors in a given cell are activated by cell surface ligands (Delta and Serrate in Drosophila) in neighboring cells in a process called ‘trans-activation’. In the absence of active ligands, Notch is in an auto-inhibited state. The binding of active ligands promotes a conformational change of the extracellular domain that renders a cleavage site accessible to metalloproteases. The resulting extracellular cleavage of Notch generates a membrane-tethered form of Notch which is the substrate of an intramembrane protease complex that cleaves Notch and releases the intracellular domain of Notch. This intracellular domain translocates to the nucleus where it associates with a DNA-binding protein to regulate the transcription of Notch target genes. Notch receptors can also interact with their ligands in cis, i.e. within the same cell, to block Notch trans-activation. This process is known as ‘cis-inhibition’.
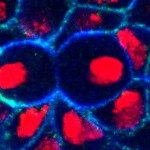
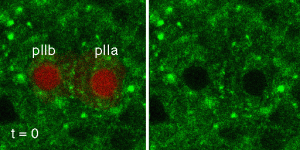
To study the dynamics of Notch signaling and trafficking in vivo, we are developing various fluorescent probes that allow us to measure the activity and trafficking of Notch in real time with single cell resolution in the living fly. In our work on Notch-mediated binary fate decision following asymmetric division, we have measured in real time the nuclear accumulation of activated GFP-tagged Notch and have shown that directional Notch signaling is rapidly established after division in one of the two Sensory Organ Precursor cell daughters (see movie below: Notch-GFP, green, chromatin of the SOP daughter cells, red).
This fate and signaling asymmetry results from the unequal segrgeation of Numb that breaks the symmetry of Notch by regulating the sorting of Notch-Sanpodo complexes at sorting endosomes (see also the project ‘Cell Polarity and Asymmetric Cell Division to learn more about Numb and asymmetric cell division). In this context, Numb appears to inhibit the recycling of Notch and promote its targeting towards late endosomes. To show this, we have developped a novel method to monitor in real time the sorting of GFP/Cherry-tagged cargoes in living cells and tissues (this made the cover of JCB). Currently, we are investigating when and where Notch is activated in this and other contexts.

L. Couturier, K. Mazouni and F. Schweisguth (2013) Numb localizes at endosomes and controls the endosomal sorting of Notch after asymmetric division in Drosophila. Current Biology, 23, 588-93
L. Couturier, M. Trylinski, K. Mazouni, L. Darnet and F. Schweisguth (2014) A fluorescent tagging approach in Drosophila reveals late endosomal trafficking of Notch and Sanpodo. The Journal of Cell Biology, 3, 351-63
M. Trylinski, K. Mazouni and F. Schweisguth (2017). Intra-lineage fate decisions involve activation of Notch receptors basal to the midbody in Drosophila sensory organ precursor cells. Current Biology, 27, 2239-47
M. Trylinski, F. Schweisguth (2019) Activation of Arp2/3 by WASp is essential for the endocytosis of Delta only during cytokinesis in Drosophila. Cell Reports, 28: 1-10
Reviews:
del Alamo, H. Rouault and F. Schweisguth (2011) Notch signalling: mechanism and functional significance of cis-inhibition. Current Biology, 21, R40-7.
F. Schweisguth (2015) Asymmetric cell division in the Drosophila bristle lineage: from the polarization of Sensory Organ Precursor cells to Notch-mediated binary fate decision. WIREs Del Biol. doi : 10.1002/wdev.175