Présentation
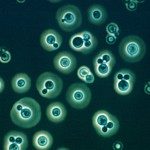
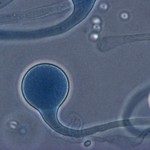
Cryptococcus neoformans and C. gattii are encapsulated yeasts that can cause life-threatening disease; mainly in immune-suppressed patients, but also in immune-competent individuals. Cryptococcosis affects the lung and may spread to the brain, manifesting itself by meningitis or brain lesions. The traversal of cryptococcus across the blood-brain barrier (BBB) is a crucial step in its pathogenesis; but why, how and when this happens is still enigmatic. Host immune cellsmost likely play a key role in this process, supported by the observation that cryptococci can survive inside macrophages after being phagocytized.
Histology and microscopy will always remain essential to investigate molecular and cellular interactions, but are limited to one observation per animal. Given that disease progression varies substantially between individual animals, the knowledge gained from histological analyses remains therefore without context for interpretation, unless it could be fit within the dynamics of the pathogenic process. In vivo imaging techniques are therefore indispensable to monitor pathogenic processes that are dynamic in time and space, to define the relevant time frames and necessary context for detailed microscopic analysis.
It is the main objective of this project to develop novel tools to dynamically monitor immune cells and cryptococci non-invasively in individual animals to resolve their interplay and interaction during the course of infection. We will image and quantify the overall progression of the infection by computed tomography (CT) and magnetic resonance imaging (MRI) and correlate it with methods that enable non-invasive and repeated imaging of targeted cells (MRI, bioluminescence (BLI), fluorescence imaging (FLI) and intravital fibered confocal fluorescence microscopy). By using multiple complementary in vivo imaging modalities, we will be able to track the different types of cells involved simultaneously and correlate this to the progression of the infection in a mouse model for pulmonary and cerebral cryptococcosis, thereby overcoming limitations of individual techniques. The in vivo results will be directly correlated with ex vivosub-cellular-resolution 3D images of whole, intact organs. Hereby, we will close the existing technological gap between in vivo imaging methods and conventional microscopy/ histology by developing optical projection tomography (OPT) and selective plane illumination microscopy (SPIM) of infected organs. Using transgenic immune cells and pathogens in combination with the here-developed multimodal imaging approach will lead to a better understanding of the dynamics of pathogenesis of cryptococcosis.
We expect that the introduction of our multi-model imaging approach will contribute to answer enigmatic key questions in infectious diseases research like how pathogens can cross physical barriers by unraveling host-pathogen interactions using dynamic monitoring of pathogenesis aspects with high resolution and sensitivity. These novel imaging and microscopy tools will be further applied for rapid in vivo screening of different (pathogenic, mutant) strains in order to identify key virulence factors, novel biomarkers, potential therapeutic targets or newdiagnostic and antifungal approaches in in vivo models of infection. Our approach is generic as it can be applied for the study of any other infection due to fungal or bacterial pathogens.
This project is headed by Uwe Himmelreich and the University of Leuven. Alexandre Alanio and Guilhem Janbon are partners.