Our main long-term goal is to develop a comprehensive methodological framework supporting the development of a quantitative understanding of cellular processes.
Our research rests on three pillars:
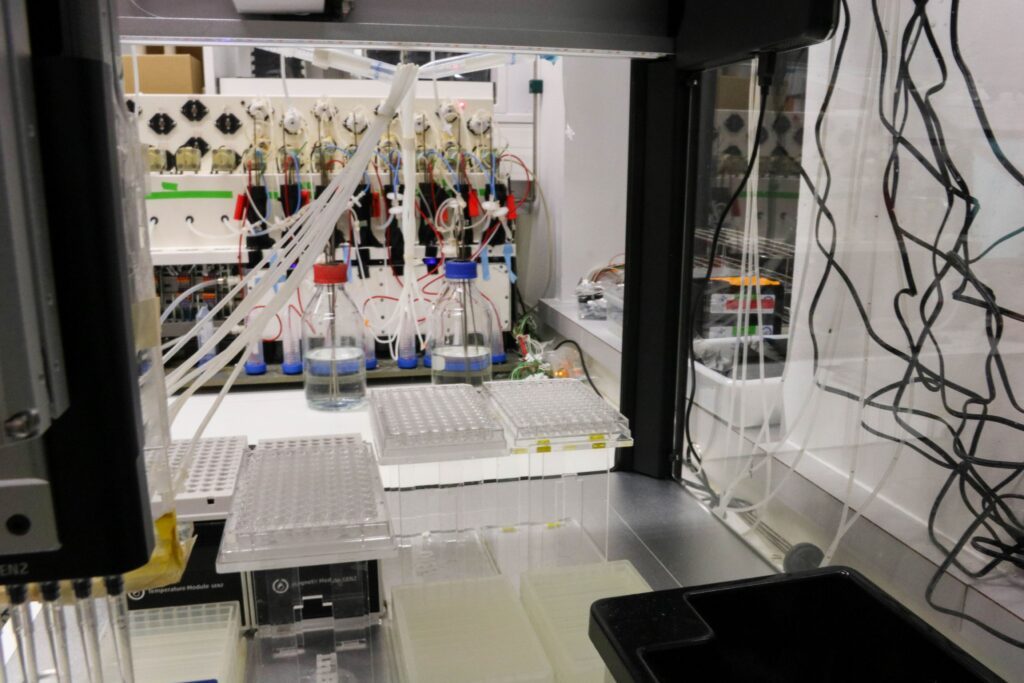
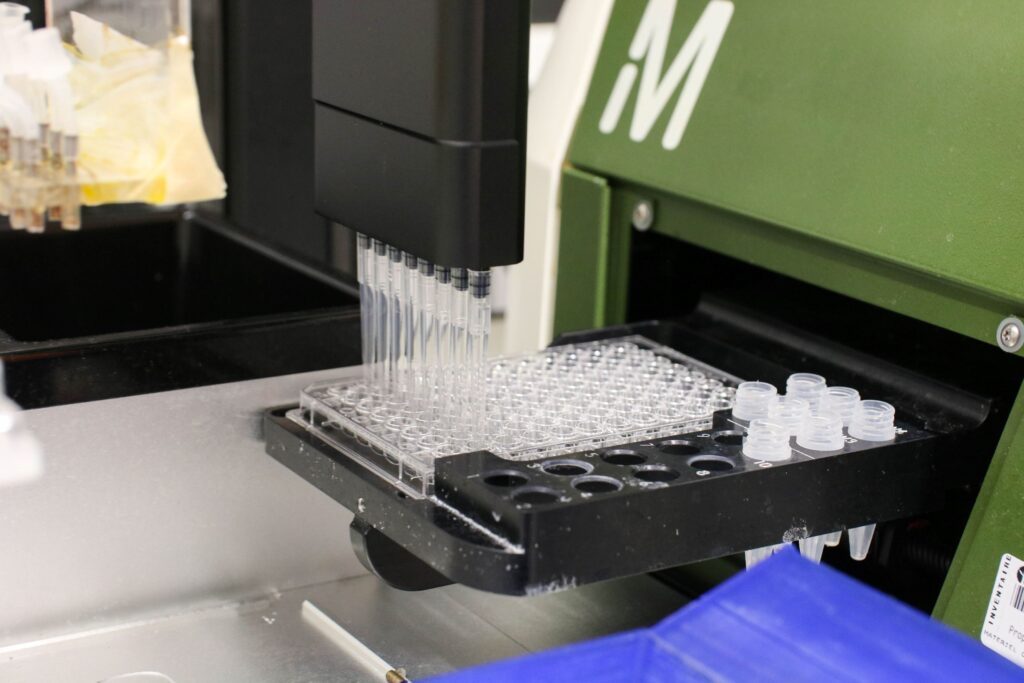
- Lab automation – engineer experimental platforms and develop computational pipeline for strain construction and data analysis,
- Bioengineering – use synthetic biology and standardized cloning approaches to probe the functioning of natural systems and construct novel systems,
- Quantitative modeling and real-time control – develop deterministic and stochastic models of cell processes to explain observations and guide designs.
Recent contributions include:
- Software supporting lab automation and real-time control of cellular processes: ReacSight (Bertaux et al, Nat Commun, 2022) and MicroMator (Fox et al, Nat Commun, 2022),
- Strategies to optimize bioproduction using regulation and control to preserve cell physiology: engineering microbial consortia (Aditya et al, Nat Commun, 2021, Aditya et al, PNAS, 2022) and applying cybergenetic principles (Sosa-Carrillo et al, Nature Commun, 2023),
- Quantifying resistance and tolerance of bacterial clinical isolates to antimicrobial treatments (Andreani et al, bioRxiv, 2021).
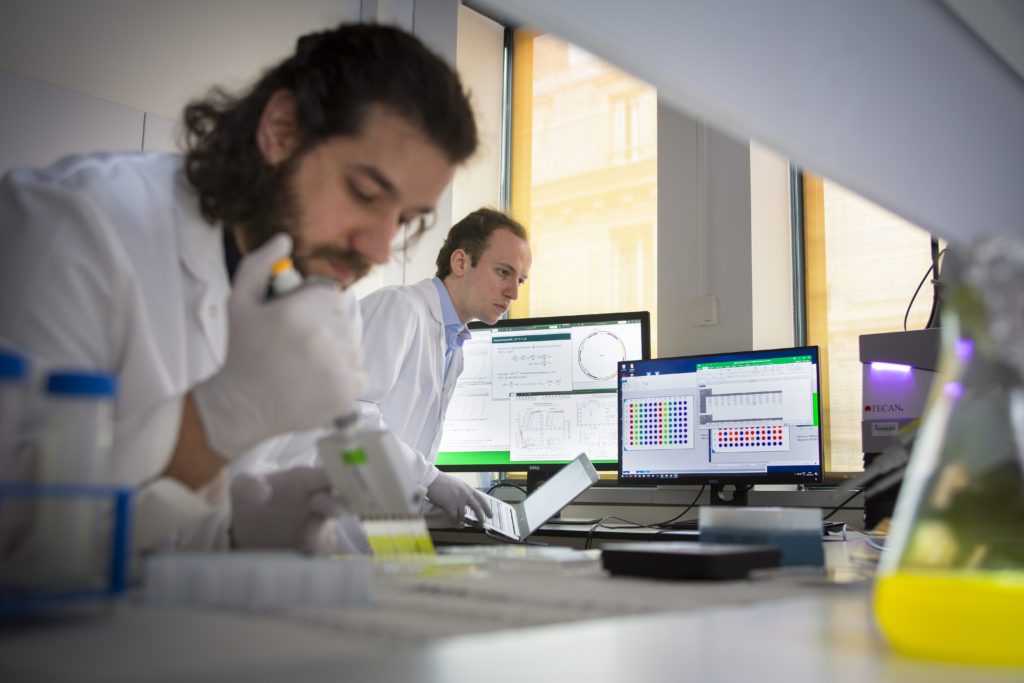
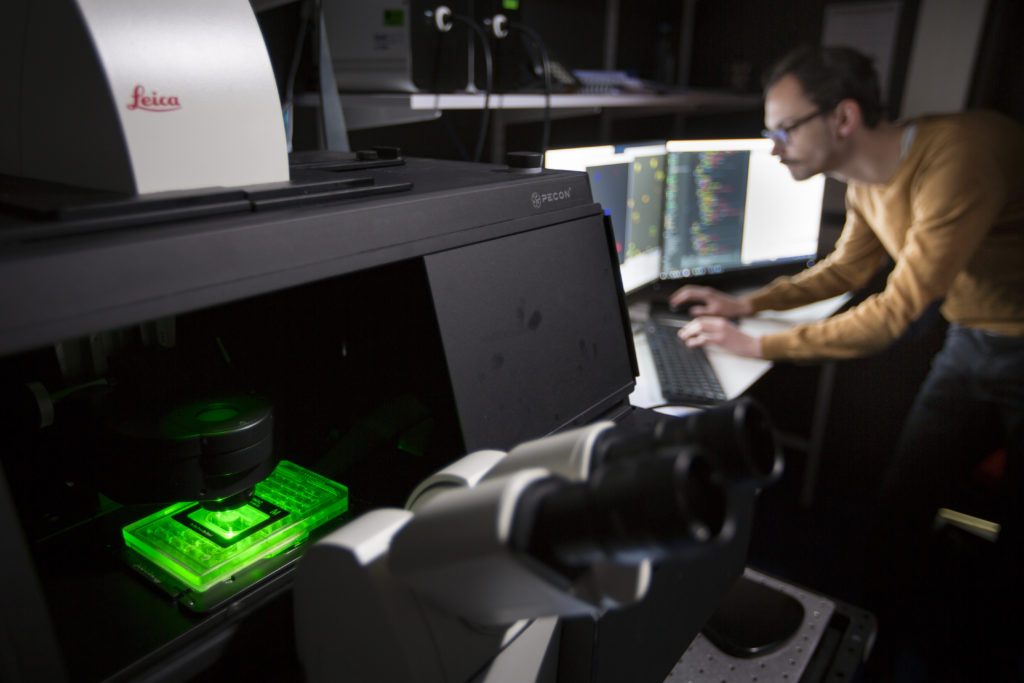
InBio is an Inria – Institut Pasteur joint research group combining wet and dry biology in the same lab. It is hosted at Institut Pasteur and affiliated to the Inria Paris research center.
New: We would like to hire a research scientist on a Inria permanent position to join our team. Inria and other French institutions offer chargé de recherche positions that are quite unique in the academic world. Several Inria researchers and engineers join forces in a small team to tackle together a grand challenge over a 10 year horizon. Our long term goal is to understand how to better engineer microbial systems by combining wet and dry biology approaches. We are located at Institut Pasteur and benefit from a truly outstanding work environment. Do not hesitate to contact us for more information.