Présentation
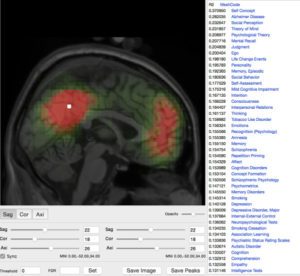
We implement and develop data analysis pipelines to process large neuroimaging-genetics datasets, and participate in the analysis of several very large international consortia. We have developed online and offline interactive tools to examine the results of many different pre-processing steps (segmentation of many regions, cortical surfaces, etc.) for hundreds or thousands of subjects.
We are also developing tools to visualise and analyse large high-resolution datasets such as those produced in 3D histology. Histological processing introduces various types of artefacts, distortions, rips, folds, etc., which had to be corrected manually. Once this first preprocessing step is finished, different automatic algorithms can be used for tissue segmentation, classification or cell-density estimation. However, the precise segmentation necessary to build a detailed brain atlas still done manually. We are developing an open, online, neuroinformatic framework to help the scientific community collaborate in the tasks that still require manual intervention, such as quality control, data annotation, and data edition – a neuroinformatic crowdsourcing framework.
Crowdsourcing has proven extremely successful in projects such as EyeWire, where online users are mapping the complete connectivity of a mouse retina, or Synapse, which allows scientists to share projects and compete to find a solution. We are developing a series of web projects exploring the utilisation of crowdsourcing for human neuroimaging, for example BrainSpell and The Brain Catalogue. We have also developed QCCC, an online crowdsourcing to quality control Freesurfer’s automatic segmentation of the corpus callosum of >1,000 subjects from the ABIDE cohort.