Présentation
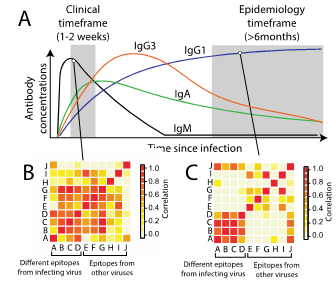
When individuals get infected by a pathogens, they will develop antibodies against that pathogen that will continue to circulate in their blood. By using assays that can detect these antibodies, we can develop an understanding of the proportion of the population that has been infected at any time. Further, by using models that integrate information about the age of the individual or information on where they reside we can provide insight into the past circulation of the pathogen. Such sero-epidemiological studies can provide a cost-efficient way of understanding past circulation of pathogens within a community and current levels of immunity.
Unfortunately, viruses from the same family may produce cross-reacting antibodies. For example, antibodies developed in response to a dengue infection will also recognize other flaviviruses, such as Japanese encephalitis, Zika or Yellow Fever viruses. Therefore, in settings where more than one flavivirus circulate, traditional serological tests can usually not allow the discrimination between different pathogens.
We are working on a number of projects in collaboration with CIBU (Dr. Jessica Vanhomwegen) that use a combination of novel serological assays that limit cross-reaction between viruses and mathematical models that properly account for the correlation structure between antibody levels to different pathogens so that we can correct for the cross-reactivity issue. These studies are performed in a number of places (including Bangladesh, French Guyana, Mali, Madagascar, Corsica (France), Burkina Faso, and Gabon) and for a number of pathogens (including dengue, Zika, chikungunya, influenza, Japanese encephalitis, Mayaro and Yellow Fever).