Présentation
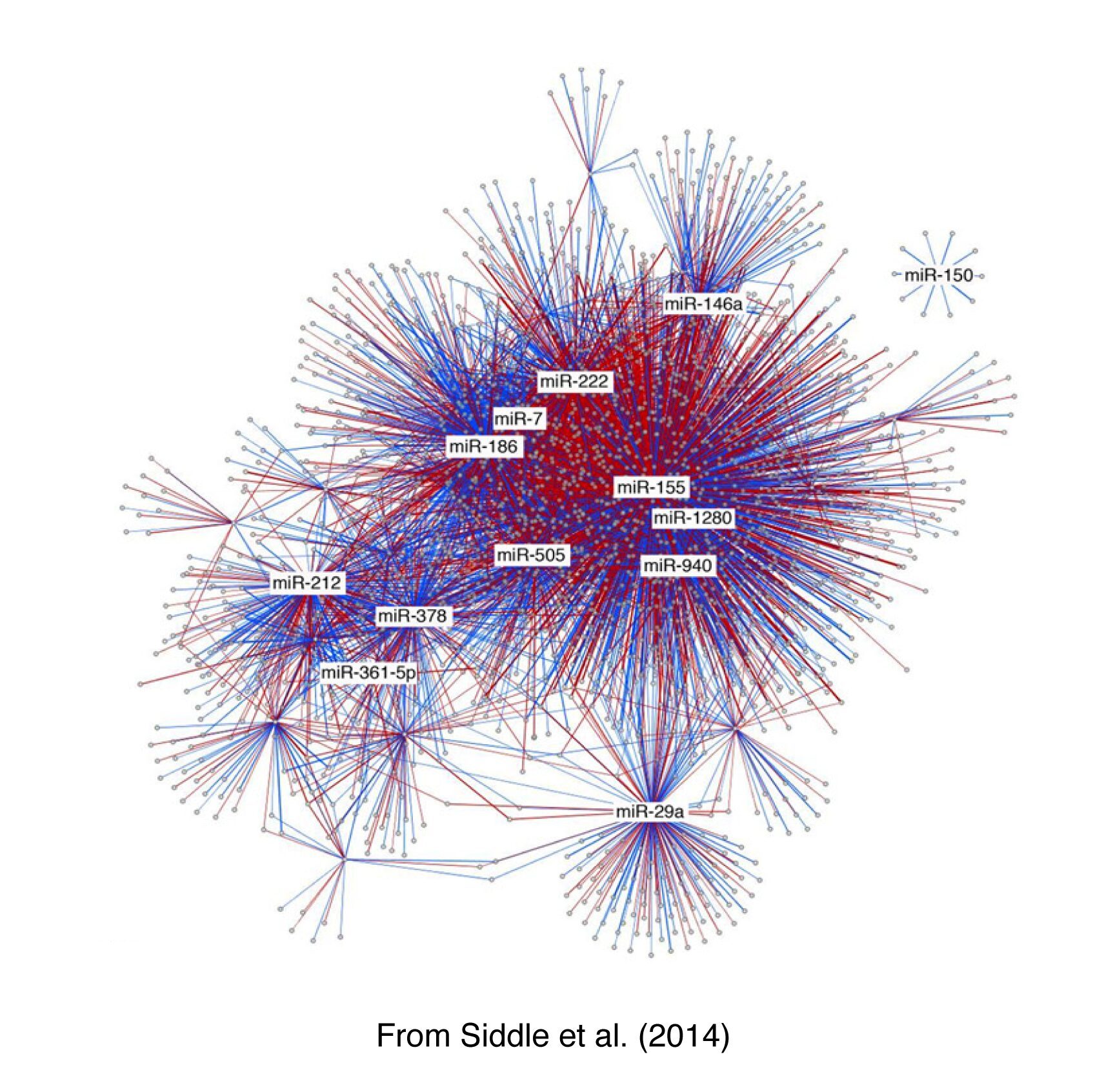
By integrating cutting-edge knowledge and technology in the fields of genomics, population genetics, immunology and bioinformatics, our aim is to establish a thorough understanding of how variable the human immune response is in the natural setting and how this phenotypic variation is under genetic control. To do so, we define population-level variation in immune responses, by studying levels of transcript abundance (of both mRNA and miRNA) before and after activation with various immune/infectious stimuli. We then map expression quantitative trait loci (eQTLs), with a particular emphasis on response/interaction eQTLs, in order to identify immune-related molecular phenotypes that have conferred a selective advantage in human adaptation. For example, we have recently assessed changes in miRNA expression upon Mycobacterium tuberculosis infection and mapped eQTL in dendritic cells. We found that the expression of 3% of miRNAs is controlled by proximate genetic factors, which are enriched in a promoter-specific histone modification associated with active transcription. Notably, we identified two infection-specific response eQTLs, providing an initial assessment of the impact of genotype-environment interactions on miRNA molecular phenotypes. We have also found that infection has a strong impact on both the relative abundance of the miRNA hairpin arms and the expression dynamics of miRNA isoforms.
In this context, our laboratory leads the EVOIMMUNOPOP project, which aims to understand how genetic variation controls transcriptional responses to TLR stimuli in populations of European and African descent. We also co-lead the MILIEU INTERIEUR Consortium (www.milieuinterieur.fr), which aims to establish the determinants of a “healthy” immune response by identifying factors (genetic, epigenetic and environmental) that contribute to the heterogeneity of immune responses in the population.