Lien vers Pubmed [PMID] – 35478717
Lien DOI – 10.1093/ve/veac029
Virus Evol 2022 ; 8(1): veac029
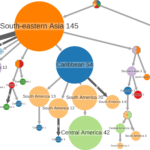
The Zika virus (ZIKV) disease caused a public health emergency of international concern that started in February 2016. The overall number of ZIKV-related cases increased until November 2016, after which it declined sharply. While the evaluation of the potential risk and impact of future arbovirus epidemics remains challenging, intensified surveillance efforts along with a scale-up of ZIKV whole-genome sequencing provide an opportunity to understand the patterns of genetic diversity, evolution, and spread of ZIKV. However, a classification system that reflects the true extent of ZIKV genetic variation is lacking. Our objective was to characterize ZIKV genetic diversity and phylodynamics, identify genomic footprints of differentiation patterns, and propose a dynamic classification system that reflects its divergence levels. We analysed a curated dataset of 762 publicly available sequences spanning the full-length coding region of ZIKV from across its geographical span and collected between 1947 and 2021. The definition of genetic groups was based on comprehensive evolutionary dynamics analyses, which included recombination and phylogenetic analyses, within- and between-group pairwise genetic distances comparison, detection of selective pressure, and clustering analyses. Evidence for potential recombination events was detected in a few sequences. However, we argue that these events are likely due to sequencing errors as proposed in previous studies. There was evidence of strong purifying selection, widespread across the genome, as also detected for other arboviruses. A total of 50 sites showed evidence of positive selection, and for a few of these sites, there was amino acid (AA) differentiation between genetic clusters. Two main genetic clusters were defined, ZA and ZB, which correspond to the already characterized ‘African’ and ‘Asian’ genotypes, respectively. Within ZB, two subgroups, ZB.1 and ZB.2, represent the Asiatic and the American (and Oceania) lineages, respectively. ZB.1 is further subdivided into ZB.1.0 (a basal Malaysia sequence sampled in the 1960s and a recent Indian sequence), ZB.1.1 (South-Eastern Asia, Southern Asia, and Micronesia sequences), and ZB.1.2 (very similar sequences from the outbreak in Singapore). ZB.2 is subdivided into ZB.2.0 (basal American sequences and the sequences from French Polynesia, the putative origin of South America introduction), ZB.2.1 (Central America), and ZB.2.2 (Caribbean and North America). This classification system does not use geographical references and is flexible to accommodate potential future lineages. It will be a helpful tool for studies that involve analyses of ZIKV genomic variation and its association with pathogenicity and serve as a starting point for the public health surveillance and response to on-going and future epidemics and to outbreaks that lead to the emergence of new variants.