Link to Pubmed [PMID] – 29273087
Link to DOI – 10.1186/s13395-017-0144-8
Skelet Muscle 2017 Dec; 7(1): 28
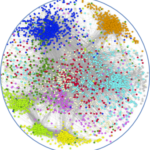
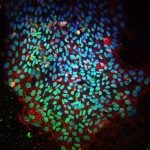
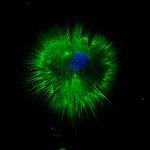
Skeletal muscle satellite (stem) cells are quiescent in adult mice and can undergo multiple rounds of proliferation and self-renewal following muscle injury. Several labs have profiled transcripts of myogenic cells during the developmental and adult myogenesis with the aim of identifying quiescent markers. Here, we focused on the quiescent cell state and generated new transcriptome profiles that include subfractionations of adult satellite cell populations, and an artificially induced prenatal quiescent state, to identify core signatures for quiescent and proliferating.Comparison of available data offered challenges related to the inherent diversity of datasets and biological conditions. We developed a standardized workflow to homogenize the normalization, filtering, and quality control steps for the analysis of gene expression profiles allowing the identification up- and down-regulated genes and the subsequent gene set enrichment analysis. To share the analytical pipeline of this work, we developed Sherpa, an interactive Shiny server that allows multi-scale comparisons for extraction of desired gene sets from the analyzed datasets. This tool is adaptable to cell populations in other contexts and tissues.A multi-scale analysis comprising eight datasets of quiescent satellite cells had 207 and 542 genes commonly up- and down-regulated, respectively. Shared up-regulated gene sets include an over-representation of the TNFα pathway via NFKβ signaling, Il6-Jak-Stat3 signaling, and the apical surface processes, while shared down-regulated gene sets exhibited an over-representation of Myc and E2F targets and genes associated to the G2M checkpoint and oxidative phosphorylation. However, virtually all datasets contained genes that are associated with activation or cell cycle entry, such as the immediate early stress response genes Fos and Jun. An empirical examination of fixed and isolated satellite cells showed that these and other genes were absent in vivo, but activated during procedural isolation of cells.Through the systematic comparison and individual analysis of diverse transcriptomic profiles, we identified genes that were consistently differentially expressed among the different datasets and shared underlying biological processes key to the quiescent cell state. Our findings provide impetus to define and distinguish transcripts associated with true in vivo quiescence from those that are first responding genes due to disruption of the stem cell niche.